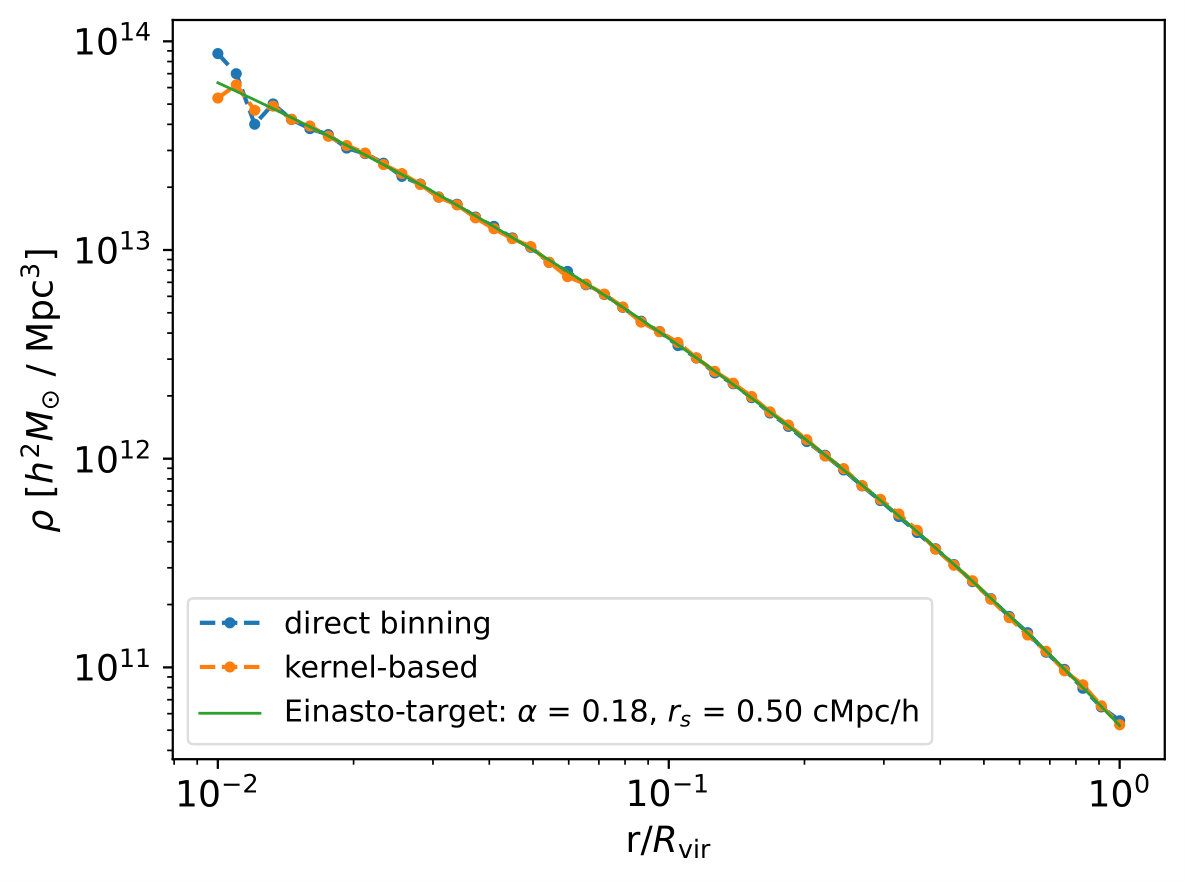
Mock Halo, Density Profile
Here, we generate 1 mock halo and calculate its mass density profile.
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Tue Mar 16 18:23:43 2021
"""
import numpy as np
import os
import subprocess
import sys
import inspect
currentdir = os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe())))
currentdir = os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe())))
subprocess.call(['python3', 'setup_compile.py', 'build_ext', '--inplace'], cwd=os.path.join(currentdir, '..', '..'))
subprocess.call(['mkdir', 'viz'], cwd=os.path.join(currentdir))
subprocess.call(['mkdir', 'cat'], cwd=os.path.join(currentdir))
sys.path.append(os.path.join(currentdir, '..', '..')) # Only needed if cosmic_profiles is not installed
from cosmic_profiles import updateCachingMaxGBs
updateCachingMaxGBs(gbs = 1)
from cosmic_profiles import genHalo, DensShapeProfs, getEinastoProf, updateInUnitSystem, updateOutUnitSystem
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams.update({'font.size': 13})
from mpi4py import MPI
comm = MPI.COMM_WORLD
rank = comm.Get_rank()
size = comm.Get_size()
def test_densities_ex_script():
#################################### Parameters ############################################################
updateInUnitSystem(length_in_cm = 'Mpc/h', mass_in_g = 'Msun/h', velocity_in_cm_per_s = 1e5, little_h = 0.6774)
updateOutUnitSystem(length_in_cm = 'kpc/h', mass_in_g = 'Msun/h', velocity_in_cm_per_s = 1e5, little_h = 0.6774)
SNAP = '017'
L_BOX = np.float32(10) # in_unit_length_in_cm
VIZ_DEST = "./cosmic_profiles/tests/viz"
CAT_DEST = "./cosmic_profiles/tests/cat"
MIN_NUMBER_PTCS = 200
CENTER = 'com'
method = {'profile': 'einasto'}
#################################### Generate 1 mock halo ##################################################
tot_mass = 10**(12) # M_sun/h
halo_res = 600000
r_s = 0.5 # Units are Mpc/h
alpha = 0.18
N_bin = 100
r_vir = np.array([1.0], dtype = np.float32) # Units are Mpc/h
a = np.logspace(-2,0.2,N_bin)*r_vir[0] # Units are Mpc/h
b = a # Units are Mpc/h. The more b differs from a, the more biased the spherically averaged density profile.
c = a # Units are Mpc/h
rho_res = 50
r_over_rvir = np.logspace(-2,0,rho_res) # Estimate density profile out to the virial radius.
model_pars = {'alpha': alpha, 'r_s': r_s}
halo_x, halo_y, halo_z, mass_dm, rho_s = genHalo(tot_mass, halo_res, model_pars, 'einasto', a, b, c)
model_pars['rho_s'] = rho_s
halo_x += L_BOX/2 # Move mock halo into the middle of the simulation box
halo_y += L_BOX/2
halo_z += L_BOX/2
print("Number of particles in the halo is {}. Mass of each DM ptc is {:.2e} M_sun/h, the average halo mass density is {:.2e} M_sun*h^2/(Mpc^3).".format(halo_x.shape[0], mass_dm, mass_dm*halo_x.shape[0]/(4/3*np.pi*r_vir[0]**3)))
dm_xyz = np.float32(np.hstack((np.reshape(halo_x, (halo_x.shape[0],1)), np.reshape(halo_y, (halo_y.shape[0],1)), np.reshape(halo_z, (halo_z.shape[0],1)))))
######################### Extract halo indices and halo sizes ##############################################
mass_array = np.ones((dm_xyz.shape[0],), dtype = np.float32)*mass_dm # In M_sun/h
idx_cat = [np.arange(len(halo_x), dtype = np.int32).tolist()]
########################### Define DensProfs object ########################################################
cprofiles = DensShapeProfs(dm_xyz, mass_array, idx_cat, r_vir, L_BOX, SNAP, VIZ_DEST, CAT_DEST, MIN_NUMBER_PTCS = MIN_NUMBER_PTCS, CENTER = CENTER)
############################## Estimate Density Profile ####################################################
# Visualize density profile: A sample output is shown above!
obj_numbers = [0]
dens_profs_db = cprofiles.estDensProfs(r_over_rvir, obj_numbers = obj_numbers, direct_binning = True) # dens_profs_db is in M_sun*h^2/kpc^3
dens_profs_kb = cprofiles.estDensProfs(r_over_rvir, obj_numbers = obj_numbers, direct_binning = False)
############################## Fit Density Profile #########################################################
r_over_rvir_fit = r_over_rvir[10:] # Do not fit innermost region since not reliable in practice. Use gravitational softening scale and / or relaxation timescale to estimate inner convergence radius.
dens_profs_db_fit = dens_profs_db[0,10:]
best_fit = cprofiles.fitDensProfs(dens_profs_db_fit.reshape((1,dens_profs_db_fit.shape[0])), r_over_rvir_fit, method = method, obj_numbers = obj_numbers) # best fit is in out units
rho_s = best_fit['rho_s']
alpha = best_fit['alpha']
r_s = best_fit['r_s']
model_pars = {'rho_s': rho_s, 'alpha': alpha, 'r_s': r_s}
cprofiles.plotDensProfs(dens_profs_db, r_over_rvir, dens_profs_db_fit.reshape((1,dens_profs_db_fit.shape[0])), r_over_rvir_fit, method = method, nb_bins = 3, obj_numbers = obj_numbers)
# Some more plotting
l_in_over_out = 1000 # Needed since in and out length units are not identical, and best_fit is in out units while r_vir is not
if rank == 0:
dens_profs_db = dens_profs_db[0]
dens_profs_kb = dens_profs_kb[0]
plt.figure()
plt.loglog(r_over_rvir, dens_profs_db, 'o--', label='direct binning', markersize = 3)
plt.loglog(r_over_rvir, dens_profs_kb, 'o--', label='kernel-based', markersize = 3)
plt.loglog(r_over_rvir, getEinastoProf(r_over_rvir*r_vir[0]*l_in_over_out, model_pars), lw = 1.0, label=r'Einasto-target: $\alpha$ = {:.2f}, $r_s$ = {:.2f} kpc/h'.format(alpha[0], r_s[0]*l_in_over_out))
plt.xlabel(r'r/$R_{\mathrm{vir}}$')
plt.ylabel(r"$\rho$ [$h^2M_{{\odot}}$ / kpc${{}}^3$]")
plt.legend(fontsize="small", loc='lower left')
plt.savefig('{}/RhoProfObj0_{}.pdf'.format(VIZ_DEST, SNAP), bbox_inches='tight')
else:
dens_profs_db = np.zeros((r_over_rvir.shape[0]), dtype = np.float32)
dens_profs_db = np.zeros((r_over_rvir.shape[0]), dtype = np.float32)
comm.Bcast(dens_profs_db, root = 0)
comm.Bcast(dens_profs_kb, root = 0)
if rank == 0:
plt.figure()
plt.loglog(r_over_rvir, dens_profs_db, 'o--', label='density profile', markersize = 4)
plt.loglog(r_over_rvir_fit, getEinastoProf(r_over_rvir_fit*r_vir[0]*l_in_over_out, model_pars), '--', color = 'r', label=r'Einasto-fit')
plt.xlabel(r'r/$R_{\mathrm{vir}}$')
plt.ylabel(r"$\rho$ [$h^2M_{{\odot}}$ / kpc${{}}^3$]")
plt.legend(fontsize="small", bbox_to_anchor=(0.95, 1), loc='upper right')
plt.savefig('{}/RhoProfFitObj0_{}.pdf'.format(VIZ_DEST, SNAP), bbox_inches='tight')